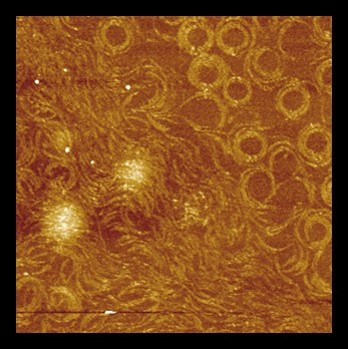
Atomic Force microscopy (AFM) *
Atomic Force Microscopy (AFM) is a technique allowing the characterization of sample morphology at nanoscale. Images are acquired by raster scanning a nanometric tip in gentle contact or intermittent contact with the sample. The tip is positioned at the end of a micrometric force transducer, the AFM cantilever: it records the variation of sample topography due to changes of tip-sample interaction during scanning operation. In addition, by measuring the tip-sample interaction force as a function of the tip-sample distance, AFMs can evaluate sample elastic and viscous properties.
AFM is nowadays recognized as an outstanding technique in the membrane field. Vertical and lateral resolution in the nanometer range can be achieved in buffer, a great advantage as compared to other structural biology techniques. In addition, due to a high signal to noise ratio, AFM often does not require image averaging, limiting the number of acquisitions. AFM has been widely used to probe at the nanoscale both topography and physical properties of biological membranes, purified or in intact cells. Besides this application, it is an instrument suitable to get high resolution pictures of filamentous structures (amyloid fibres, DNA, intrinsically disordered proteins…) as well as viral particles or bacteria.
Initially limited by the time to acquire an image, tremendous progress in AFM during the last 10 years laid on the development of high-speed (HS)-AFM that enables dynamic imaging of biological samples.
In addition, AFM topography can also be combined with fluorescence microscopy (conventional and super resolution microscopy as well as spectroscopy) to correlate topography and mapping of a component of interest.