EVOLVE Distributed In-Person Training Course: Intro to BioImaging Analysis with Python for Life Scientists
Event details
- When
- June 23, 2026 09:00–June 25, 2026 17:00 CEST
- Where
- Czech Republic - Portugual - Sweden - UK

Interested in learning how to use Python for your image analysis needs? Want to understand how Jupyter notebooks can make your analysis workflows easier?
The second edition of the distributed "Image analysis with Python" beginners course is your chance. The course will take place June 23rd - 25th 2026. As a distributed course, it will be held in-person in four locations, with a joint training program, common lectures and in-person hands-on support.
With support from the Euro-BioImaging team as part of the EVOLVE project (GA#101130986), the course will be held on site across different Euro-BioImaging Nodes at the following locations:
- University of Gothenburg, Sweden - NMI Sweden Node
- Gulbenkian Institute for Molecular Medicine (GIMM), Oeiras, Portugal - PPBI Node (with the collaboration of ITQB - NOVA )
- The Francis Crick Institute, London - UK Node
- BIOCEV, Vestec, Czech Republic - Prague Node
Prerequisites for participation
- Working with biological imaging techniques and having a clear image analysis question. You have to submit short paragraph describing your image analysis challenges.
- Basic knowledge of ImageJ (opening datasets, basic processing techniques such as filtering and segmentation, object quantification such as “Analyze Particles”). Some exposure to coding (e.g., ImageJ macro writing) would be beneficial.
- Students must be able to attend the whole 3-day course, no exceptions.
- Please bring a laptop with a minimum of 8GB RAM, that you can connect to WiFi, with at least 14-16” screen + mouse (not just mousepad) for the course.
What’s included in the course?
This distributed course will be held in person at the four locations and the four local training hubs will be remotely connected and jointly going through the training programme provided each day by one of the sites.
The course will implement BAND as a backup joint virtual platform for students unable to meet course working environment requirements. This virtual platform is the BAND remote desktop, developed as part of the Euro-BioImaging involvement in the EOSC-Life project by Jean-Karim Heriche and Yi Sun from EMBL [link]. This training event also benefits from the EGI Training Infrastructure service supported by national resource providers in the EGI Federation. In particular, CESNET is providing the computational resources and support to host BAND for this training event.
Venues and teams providing the training
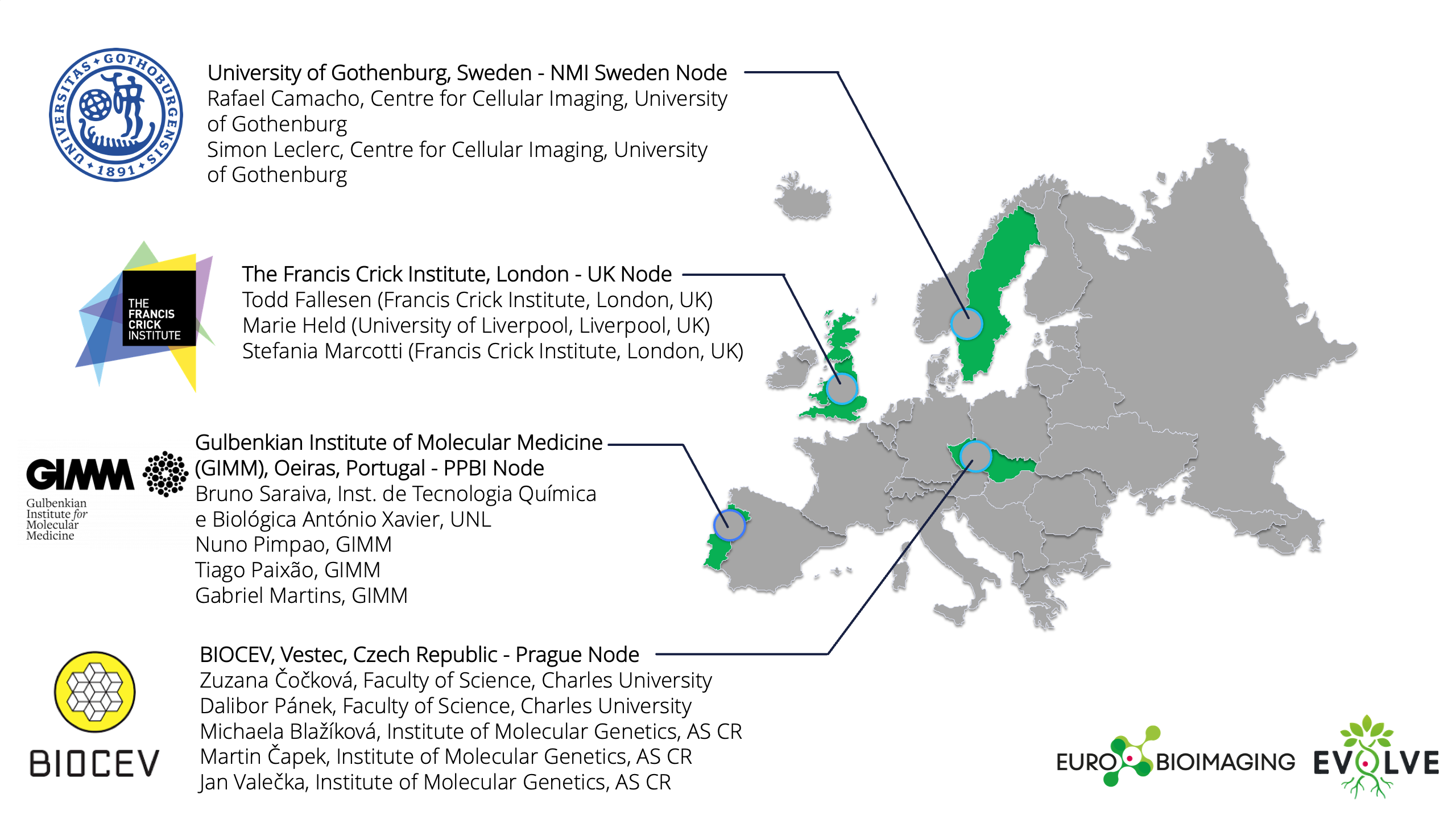
Registration is open until April 15th , and local travel grants are available here.
Program
Day 1: Intro to Jupyter notebooks and python, introduction to loading images and microscopy metadata, basics of visualization (Sweden Node).
Day 2: Practical image processing with Python: segmentation, denoising, morphology and automation of analysis pipelines (UK Node)
Day 3: Novel strategies for segmentation: Machine learning + use of pre-trained models from BioImage Model Zoo (Portuguese + Prague Nodes).
| Time in CEST | Tuesday, Jun 23rd | Wednesday, Jun 24th | Thursday, Jun 25th |
|---|---|---|---|
| 9:30-10:00 | Welcome/Arrival | Optional Q&A for CEST timezone locations | Optional Q&A for CEST timezone locations |
| 10:00-11:30 | Introduction to Jupyter and python for the context of bioimage analysis | Basics: Handling images in Python | Introduction to Artificial Intelligence and classical Machine Learning segmentation |
| 11:30-11:45 | Coffee break | Coffee break | Coffee break |
| 11:45-13:00 | How to create our own kernel | Denoising and segmentation | Machine Learning classification tasks |
| 13:00-14:00 | Lunch | Lunch | Lunch |
| 14:00-15:30 | Introduction to microscopy images – images as scientific dataI/O | Feature Extraction | Introduction to Neural Networks and Deep Learning |
| 15:30-15:45 | Coffee break | Coffee break | Coffee break |
| 15:45-17:00 | Bioimage analysis workflow – example as an introduction to the rest of the course | Building a workflow/pipeline | Accessible tools to use DL for image analysis |
| 17:00-17:30 | Optional Q&A for BST timezone locations | Optional Q&A for BST timezone locations | Optional Q&A for BST timezone locations |