
June 3, 2026
BMIC meets Flanders BioImaging
In May 2026, the Belgian Molecular Imaging Community (BMIC) and Flanders BioImaging joined forces in Ghent for a two-day community event that brought…
Katrín Möller is a post-doctoral researcher at Biomedical Center of the University of Iceland, where she studies epigenetics and neuronal development. She’s just published a paper in eLIFE entitled “A role for the centrosome in regulating the rate of neuronal efferocytosis by microglia in vivo,” which provides important insight into microglial phagocytosis and features impressive light sheet microscopy images. Beyond describing exciting science in her publication, Katrin Möller also uploaded the underlying bioimaging data to the Bioimage Archive. Here she outlines some of the reasons for depositing her epic dataset and highlights a few benefits of data sharing for researchers.
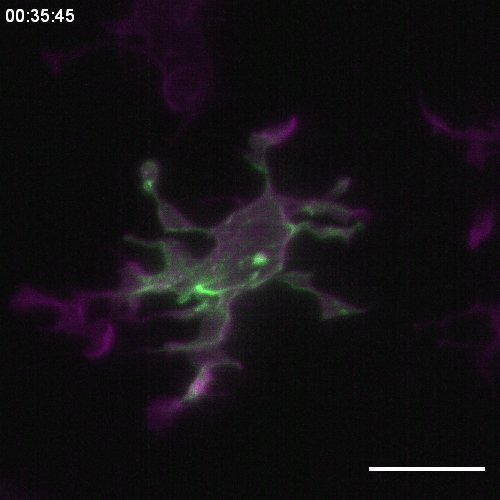
“The goal of our study was to better understand how microglia behave in the developing brain, in particular during the microglia engulfment process. This is a subject that has been studied on a broad scale, but we wanted to look at it on the intracellular and molecular level. We used Zebrafish embryos as a model system as they are optically transparent, thereby allowing live imaging of fluorescently labelled microglia in their native environment, the brain. We used this setup to examine not only how the microglia behave collectively, but also what’s going on inside them,” explains Katrín.
“We chose light-sheet microscopy to capture the dynamic microglia behaviour because it is very fast but also much gentler for live cells and full organisms than conventional confocal microscopy. The flipside is that it creates massive amounts of data. Over the course of this project, almost 44 Terabytes of data were created, which became ~700 Gigabytes after curation.”
Katrín carried out the imaging work during her PhD at the microscopy core facility of the University of Zurich. The facility provides server space where the imaging data can be stored for a few months for post-processing, before it is transferred to the research groups own servers for long-term storage. “The facility also provides high-performance virtual computers for image data processing,” adds Katrín. “Thank goodness, because my own personal computer wasn’t nearly powerful enough to process that type of dataset!”
Benefits of sharing data
As she analyzed her data, she realized she had captured some very unique views of intracellular components – including rare centrosome movements. These displacements are important for the phagocytic process and potentially demonstrate something common for immune cells, and could thus be of great interest to the wider scientific community. Katrín began to consider making her dataset FAIR (Findable, Accessible, Interoperable and Reusable – see reference 1 below), so other scientists could reproduce the study and reuse her dataset for future projects.
“I was really motivated to share this data. It’s a first step towards saving resources. I would of course be very excited if we could learn something new about microglia using my data. But my curated data set also contains for example 40-50 zebrafish brains. If a scientist uses this dataset first, before acquiring their own data, it could potentially save them a step. It could give them a starting point to test their pipelines, or allow them to compare behavior of zebrafish microglia to another system. In addition, it could be useful for people who are developing software or training an algorithm.”
But it was the eLife journal’s requirement that all related data should be stored in a public repository that convinced Katrín to follow through on her idea. The next step was to choose a public repository.
BioImage Archive – a home for life-sciences microscopy data
“I was wondering where to store my data when I learned about the BioImage Archive at the BNMI Symposium from Euro-BioImaging project manager Johanna Bischof. It’s great to have a place to put life-sciences microscopy data.”
Supported by the Euro-BioImaging and BioImage Archive team, Katrín went the extra mile, and submitted all of the imaging datasets behind the figures in her paper: a total of 700 Gigabytes of data in ~600 files. “Afterall, the more of your data you share, the more believable your study is,” adds Katrín. “And a final benefit of sharing the data for our lab is that we now have a secure, long-term place to house the dataset which frees up active storage space for new experiments.”
View the data: https://www.ebi.ac.uk/biostudies/BioImages/studies/S-BIAD564
Submitting data to the BioImage Archive
“I have to say that the study was accepted really fast by the journal; it took less than 1 month,” says Katrín. “But it took me an additional month to prepare the data for submission to the BioImage Archive.” The data had to be categorized based on the type of experiment, links between the data and analysis files had to be prepared and each file had to be annotated. “Especially finding all the metadata was very time-consuming. In my case, the most important metadata were the zebrafish lines and organelle markers, and the image resolution (spatial and temporal), but in general, anything that helps a person who looks at your data to understand what they’re looking at is important to share,” explains Katrín.
As Katrín learned, it is important to start submitting the data to the BioImage Archive early on – at the same time as you submit your paper. “I would recommend starting immediately - because it will take time to prepare and upload everything,” says Katrín. “And you don’t have to worry about your data being online too early. You can tell the BioImage Archive team to not make the data public until the paper is published.”
“The BioImage Archive really is a fantastic effort. Despite the hard work, I’m really glad I did this. It feels like a big step towards more sustainable science,” concludes Katrín.
Read the full article:
Katrin Möller, Max Brambach, Ambra Villani, Elisa Gallo, Darren Gilmour, Francesca Peri (2022) A role for the centrosome in regulating the rate of neuronal efferocytosis by microglia in vivo eLife 11:e82094
https://doi.org/10.7554/eLife.82094
About the BioImage Archive
The BioImage Archive (see reference 2 below), hosted by EMBL-EBI, is a free of charge, publicly available online resource which stores and distributes biological images. It accepts submissions of data from any imaging modality, for which no other, more specialized repository exists as long as the data are either associated with a peer-reviewed publication, or of value beyond a single experiment. The BioImage Archive supports and promotes FAIR data sharing and implements the REMBI (‘recommended metadata for biological images’ - reference 3 below) metadata guidelines to enable FAIR data.
References:
1) Wilkinson MD, Dumontier M, Aalbersberg IJ, et al. The FAIR Guiding Principles for scientific data management and stewardship [published correction appears in Sci Data. 2019 Mar 19;6(1):6]. Sci Data. 2016;3:160018. Published 2016 Mar 15. doi:10.1038/sdata.2016.18
2) Hartley M, Kleywegt GJ, Patwardhan A, Sarkans U, Swedlow JR, Brazma A. The BioImage Archive - Building a Home for Life-Sciences Microscopy Data. J Mol Biol. 2022;434(11):167505. doi:10.1016/j.jmb.2022.167505
3) Sarkans U, Chiu W, Collinson L, et al. REMBI: Recommended Metadata for Biological Images-enabling reuse of microscopy data in biology. Nat Methods. 2021;18(12):1418-1422. doi:10.1038/s41592-021-01166-8

June 3, 2026
In May 2026, the Belgian Molecular Imaging Community (BMIC) and Flanders BioImaging joined forces in Ghent for a two-day community event that brought…

June 1, 2026
The EOSC Association has adopted a new position paper calling for a distinct Work Programme-based Partnership for EOSC in FP10. Euro-BioImaging welcomes this…
May 29, 2026
Spatial transcriptomics is rapidly transforming the way researchers study complex biological systems. To explore the latest developments in this fast-moving field, Euro-BioImaging is…