April 29, 2026
Welcome to our new Smart Optical Microscopy (SOM) Node!
We are delighted to announce that a new Node has joined Euro-BioImaging after stringent review by our Scientific Advisory Board and approval by the…
Meiofauna are tiny marine organisms that range in size from 20 microns to 1 milimetre. They are present everywhere in the sea, from the shore to the deep ocean. They are an important part of the food web and a crucial link between micro- and macro- ecosystems. Nevertheless, to this day, very little is known about these creatures. Valentin Foulon, a research engineer in marine biology, is part of the Blue Revolution program that aims to develop a taxonomic identification protocol for meiofauna. But meiofauna are hard to observe – too large to observe with traditional microscopes and yet too small to see with the naked eye. So, Valentin contacted the Bretagne-Loire node of France-BioImaging, in Nantes, where a range of high-end microscopes are available in open access. We spoke to Valentin to find out if Single Plane Illumination Microscopy (SPIM) light-sheet microscopy may literally shed light on this exciting new frontier in ocean sciences.
Valentin Foulon loves imaging and microscopy – and he loves meiofauna. He works on the Blue Revolution program, a nationally-funded project that brings together three institutions in Brittany region of France: French National Institute for Ocean Science (Ifremer), École Nationale d'Ingénieurs de Brest (ENIB) and the Station Biologique de Roscoff. The goal of the project is to accelerate the taxonomic identification of meiofauna using artificial intelligence algorithms. The project started in 2020, to capitalize on a taxonomic identification protocol that was developed for plankton during the TARA Ocean expedition. But while the confocal-microscopy-based protocol worked well for small plankton, the researchers quickly realized that it had to be adapted to work with meiofauna.
Choosing the right microscopy approach
“Identification of meiofauna is incredibly challenging,” says Valentin Foulon. “To identify these organisms, we first have to sort from the sediment in which they live. This is done manually. Then we mount them on the microscope slide, and look closely at small details to describe the genus,” explains Valentin. “It’s very labor-intensive. We have to be patient because it takes a lot of time to separate the meiofauna from organic matter. “On a classical microscope, we have only one focal plane. To see important details, we need a 3D view of organisms. Confocal microscopy allows this by making multiple optical slices. However, confocal microscopy was not adapted to the big size range of meiofauna.” explains Valentin. “That’s why we got curious about light-sheet imaging, because we would be able to see the organisms in 3D.”
While light-sheet microscopy seemed technically promising, there was no light-sheet microscope available for Valentin and his team to use in all of Brittany. “But there was one in Nantes, which isn’t far from Brest,” says Valentin. “And it is available to use in open access with France BioImaging. We clearly saw an opportunity there.”
Valentin’s project proposal, which was submitted to the Euro-BioImaging pilot User Access fund, was fully funded by France-BioImaging, in a generous initiative introduced by France-BioImaging to fund all user projects to its facility received by Euro-BioImaging for the pilot User Access fund.
“I had never tried light sheet microscopy before,” says Valentin, “And in Nantes, the lab didn’t have experience with marine organisms. So before running the experiment, I went to Nantes for 3 days, to test the set-up and prepare the samples with them, to ensure the protocol worked before running it. We mounted the organisms in agar capillary. Luckily, we managed to have a good staining in order to see what we wanted to see. Together, we found a compromise between the resolution, the acquisition time, etc. Once the set-up was fine-tuned, I sent my samples, and the engineer at the lab did the imaging work.”
“It was a very nice experience,” says Valentin. “I discovered a new technique, which is always enriching. I also think the lab was happy to work with a new sample type. And finally, we validated our proof-of-concept, proving that it is possible to image these organisms in 3D.”
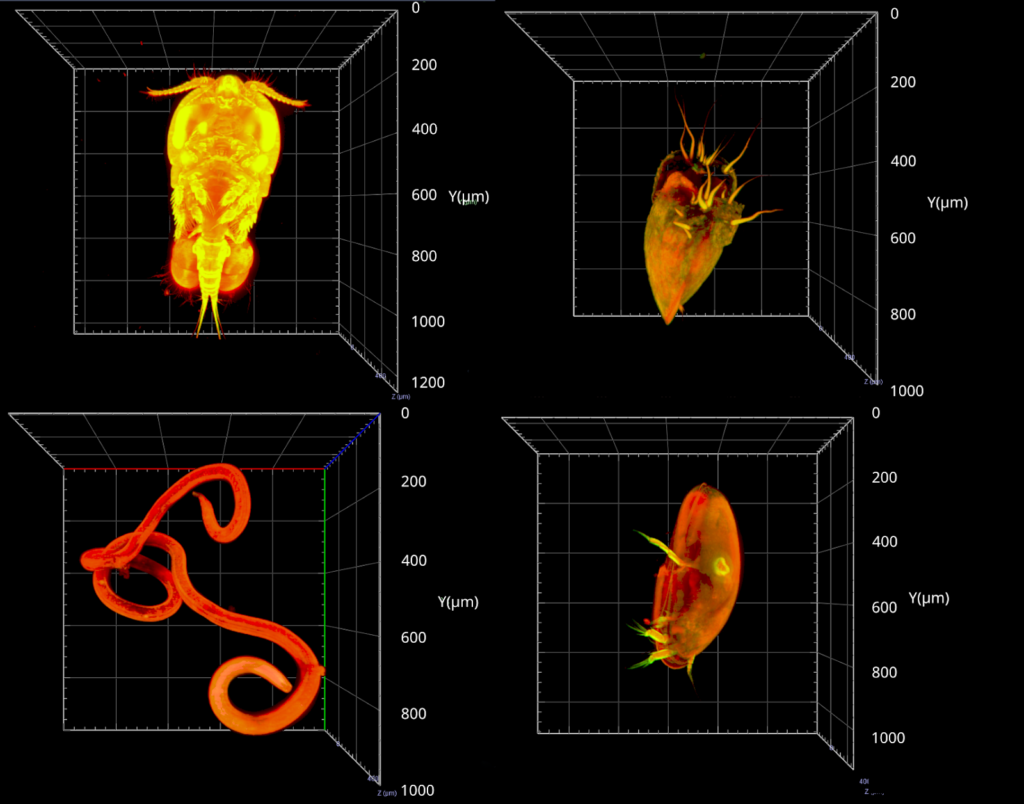
Image data management
But the project doesn’t end there. “The next step is data management. We imaged between 200-300 different organisms. It’s not enough for automatic classification, but nevertheless, we have 6-7 TB of data. Now we must process the data for automatic classification with machine learning. It’s an essential part of the project, to make the link between sample imaging, and data processing. We are working to close that gap.”
“The outcome of this project is to validate a proof-of-concept, from the sample collection to the image classification. With this new taxonomic identification protocol, we will be one step closer to understanding meieofauna, these mysterious marine organisms, who may hold the key to understanding the impact of human activity on marine ecosystems,” concludes Valentin.
About France BioImaging
The France-BioImaging Node is at the crossroads between molecular and cell biology, integrated physiology, biophysics and engineering, mathematics and informatics. This unique coordinated infrastructure in the European landscape gathers several large facilities and R&D laboratories specializing in imaging in 6 National Geographic locations and one transversal Node in bioimage informatics. FBI aims at creating the most efficient adoption of the latest advances in technologies and methods related to microscopy, by the users of the imaging facilities. R&D labs agree to open their technologies and expertise to the European community either by hosting users on their own site or after technological transfer to the FBI core facilities. These technologies and methods, reinforced by a strong support in computational analysis, provide quantitative measures and integrative understanding of a wide range of cell and tissue activities in biological models, from the simplest organism, to small animals in normal and pathological situations.
April 29, 2026
We are delighted to announce that a new Node has joined Euro-BioImaging after stringent review by our Scientific Advisory Board and approval by the…

April 29, 2026
A new publication from the Global BioImaging community sheds light on the career paths, recognition, and working conditions of imaging…
April 28, 2026
AgroServ is a transdisciplinary initiative supported by the European Union through the Horizon Europe program and will continue until 2027. It supports the research…