March 26, 2026
Welcome, MAdMiC Node!
The Madrid Advanced Microscopy Center (MAdMiC) is the first Euro-BioImaging Node in Madrid (Spain). It is formed through the collaboration and close work of…
Cell division is the fundamental process in living organisms - the basis for growth and survival of both single- and multicellular systems. As part of cell division, the duplicated DNA, in the form of mitotic chromosomes, has to be distributed across the two daughter cells, with each receiving exactly one copy of each chromosome. Mistakes in chromosome segregation will result in cell death, cancer, or severe developmental diseases.
Truly understanding the process of chromosome segregation in detail has been a struggle - the process is fast, three-dimensional, and happening in a very condensed space, challenging different characterisation methods.
To address this challenge, the team of Stoyno Stoynov from the Institute of Molecular Biology, Bulgarian Academy of Sciences, Sofia, and from our Advanced Light Microscopy Sofia Node, introduce Facilitated Segmentation and Tracking of Chromosomes in Mitosis Pipeline (FAST CHIMP). This method brings together time-lapse super-resolution microscopy in the form of Airyscan imaging with deep learning to allow tracking of chromosome dynamics at high spatiotemporal resolution.
These two methods are uniquely suited to address two challenges in following chromosomes during cell division. Airyscan super-resolution imaging is fast enough while comparatively low in light intensity, reducing photobleaching and phototoxicity. The team developed and combined deep learning methods for denoising, segmentation, and registration, specifically developed for the chromosome dataset. Chromosomes during mitosis are difficult to segment and track automatically due to their uneven and varying shape, changing volume, and their crowding.
The team tested the developed tools in a range of different cells and studied chromosome location and movement both during normal as well as abnormal cell division processes. They observed that chromosomes during mitosis move according to a flow field, indicating substantial rearrangements of their position and opportunities for translocations across chromosomes. FAST CHIMP can also track both intra- and interchromosomal contacts in high resolution and over longer time intervals.
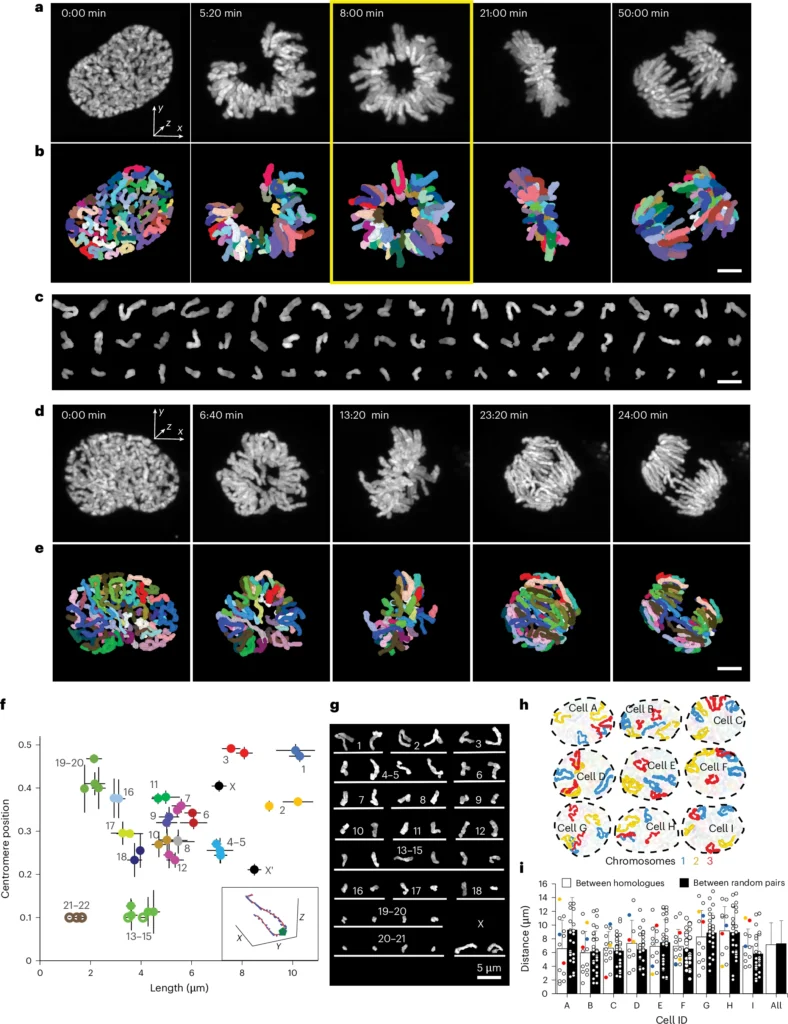
The new deep learning methods provide an important step forward, but they continue to rely on high-frequency super-resolution imaging, which is a technique that requires both specialised instrument setups as well as specific expertise, such as that available at the facility in our Sofia Node.
The excellent work is published in Nature Cell Biology, including a prominent spot on the cover of the April edition.
Read more in: Stamatov, R., Uzunova, S., Kicheva, Y. et al. Supra-second tracking and live-cell karyotyping reveal principles of mitotic chromosome dynamics. Nat Cell Biol 27, 654–667 (2025). https://doi.org/10.1038/s41556-025-01637-6
Congratulations to the team for this breakthrough development!
You can also hear from first author Rumen Stamatov in our Virtual Pub seminar on June 13th - Learn more.


March 26, 2026
The Madrid Advanced Microscopy Center (MAdMiC) is the first Euro-BioImaging Node in Madrid (Spain). It is formed through the collaboration and close work of…

March 26, 2026
German BioImaging, within its work in the NFDI4BIOIMAGE consortium, and in collaboration with Euro-BioImaging ERIC, has launched a new survey to collect input about…

March 25, 2026
Turin, Italy – 20–22 October 2026 Early career professionals working in imaging core facilities will soon have the opportunity to strengthen essential skills beyond…