
June 3, 2026
BMIC meets Flanders BioImaging
In May 2026, the Belgian Molecular Imaging Community (BMIC) and Flanders BioImaging joined forces in Ghent for a two-day community event that brought…
Today, imaging scientists are producing datasets of increasing volume, complexity, and information content. To realize the full potential of imaging data, Euro-BioImaging Nodes offer a range of Image Data Services, including data handling, developing bespoke image analysis workflows, quantitative analysis, and access to analysis software. Access to these image analysis services as a standalone service is currently available at selected Nodes as part of our Proof-of-Concept study. We spoke with Jean-Yves Tinevez of the Image Analysis Hub (IAH) at Institut Pasteur, part of France BioImaging Node, to learn more about how the image data services his team offers can help biologists get the most out of their data.
What type of image data services do you and your team offer?
Jean-Yves Tinevez: At the Image Analysis Hub at Institut Pasteur, we develop custom bioimage analysis tools and pipelines, mainly focusing on live-cell 2D or 3D over time images, mainly coming from optical microscopy. We work by collaborating scientifically with our users, trying to achieve a significant added-value to their Research. We offer two types of services:
1. Quantification projects. In this scenario, a user would delegate an analysis task to us, a sub-part of their research project. They transfer the image dataset to us and we perform the analysis. Often, we enter a project before images are acquired, and we can work with our colleagues from the imaging facility on experimental design. Our goal then is to produce scientific results from their images, up to the production of a figure for a publication, with the redaction of the associated Materials and Methods section.
2. Custom development of analysis tools. With this type of project, we develop, validate and deploy a bespoke analysis pipeline for a user. The analysis is then performed by the user. We strive not to limit ourselves to be users of bioimage analysis software, but to be able to build new tools. Development can take very long. To accelerate the project, we base our development on existing scientific software platforms (such as Fiji, Icy, QuPath ...) or the software we develop and maintain (among which TrackMate or related tools like MaMuT and Mastodon). We use our expertise to help other scientists get the most information out of their images.
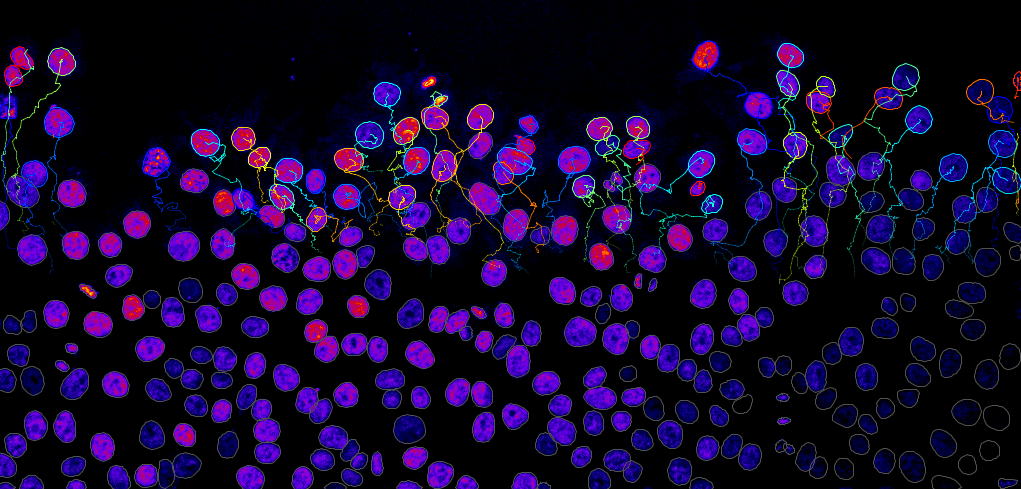
What is a typical project like?
Jean-Yves Tinevez: During a project, engineers from our facility collaborate with users and create a tool, adapt a pipeline or perform an analysis in order to answer a scientific question. This involves scientific work from us beyond a purely technical contribution. Some of our collaborative projects have been published in peer-reviewed journals. Sometimes our engineers are included in the author list, acknowledging the scientific responsibility and commitment to the project. This is the type of project we want to develop further – where we can truly help researchers overcome a bottleneck in their work. We also have significant experience in providing services towards such projects.
We serve on average 110 users per year, of which 9% are external to the Institut Pasteur. We accept 20 to 40 new projects every year, and a project can involve from 10 to 200 hours of effort, spread over weeks to years.
How are the projects selected?
Jean-Yves Tinevez: There is no selection committee for IAH project submissions, but there is a feasibility assessment, and the time spent on a project is billed to the user. Feasibility assessment includes cost and time considerations, as well as image quality vs desired robustness of results. If a project is deemed feasible, the proposal is accepted and treated in order of submission.
How can we get started?
Anyone interested in trying the Image Data Analysis services offered by France BioImaging can apply via the Euro-BioImaging portal. Or you can join the open-desk sessions we organize with France-BioImaging to get started with a consultation. And for more information, please contact us!
Interview with Jean-Yves Tinevez on YouTube.
REFERENCES:
Ershov, Dmitry, Minh-Son Phan, Joanna W. Pylvänäinen, Stéphane U. Rigaud, Laure Le Blanc, Arthur Charles-Orszag, James R. W. Conway, et al. “Bringing TrackMate into the Era of Machine-Learning and Deep-Learning.” bioRxiv, September 20, 2021. https://doi.org/10.1101/2021.09.03.458852.
Tinevez, Jean-Yves, Nick Perry, Johannes Schindelin, Genevieve M. Hoopes, Gregory D. Reynolds, Emmanuel Laplantine, Sebastian Y. Bednarek, Spencer L. Shorte, and Kevin W. Eliceiri. “TrackMate: An Open and Extensible Platform for Single-Particle Tracking.” Methods, Image Processing for Biologists, 115 (February 15, 2017): 80–90. https://doi.org/10.1016/j.ymeth.2016.09.016.
Wolff, Carsten, Jean-Yves Tinevez, Tobias Pietzsch, Evangelia Stamataki, Benjamin Harich, Léo Guignard, Stephan Preibisch, et al. “Multi-View Light-Sheet Imaging and Tracking with the MaMuT Software Reveals the Cell Lineage of a Direct Developing Arthropod Limb.” Edited by Alejandro Sánchez Alvarado. ELife 7 (March 29, 2018): e34410. https://doi.org/10.7554/eLife.34410.
Mastodon: https://github.com/mastodon-sc/mastodon

June 3, 2026
In May 2026, the Belgian Molecular Imaging Community (BMIC) and Flanders BioImaging joined forces in Ghent for a two-day community event that brought…

June 1, 2026
The EOSC Association has adopted a new position paper calling for a distinct Work Programme-based Partnership for EOSC in FP10. Euro-BioImaging welcomes this…
May 29, 2026
Spatial transcriptomics is rapidly transforming the way researchers study complex biological systems. To explore the latest developments in this fast-moving field, Euro-BioImaging is…